Mosquitoes and other blood-feeding insects transmit many important viral pathogens. Some of these arthropod-borne (arbo-)viruses are responsible for worldwide epidemics and high mortality rates. For example, Dengue virus, a positive-sense (+) RNA virus from the Flavivirus family, causes an estimated 50-100 million infections per year worldwide and can causes severe, sometimes fatal disease. Arboviruses replicate efficiently in both mosquito vector as well as in its vertebrate host. These viruses thus encounter the innate and adaptive immune responses of vertebrates, as well as the — poorly characterized — innate immune system of insects. We study virus-host interactions in insects and mammals, with a major focus on RNA silencing pathways, innate immunity and the identification of antiviral compounds.
Specific interests include:
- Antiviral RNAi.
RNA interference (RNAi) — a mechanism for gene silencing guided by small interfering RNA (siRNA) — is an important antiviral immune response in the fruitfly Drosophila melanogaster and in mosquitoes. In this pathway viral double-stranded RNA (dsRNA) is recognized as non-self and processed into virus-derived siRNA (vsiRNAs). These vsiRNAs are incorporated into Argonaute-2, the central component of the RISC complex. Argonaute 2 then uses the vsiRNAs to find viral RNAs to induce their degradation. The RNAi pathway thus restricts viral replication at two levels: by cleaving viral dsRNA molecules replication intermediates and by destroying single-stranded viral RNAs.
As a counter-defense to the powerful antiviral RNAi response, viruses have evolved sophisticated mechanisms to suppress or evade it. We have recently identified virus-encoded suppressor proteins in mosquito- and Drosophila viruses. These proteins interfere with different steps of the RNAi pathway. For example, the Nora virus VP1 protein suppresses the catalytic activity of Argonaute-2, underlining the importance of Argonaute 2 in antiviral defense. Several other viral suppressors of RNAi that we recently identified bind dsRNA to interfere with vsiRNA biogenesis and to prevent their incorporation into RISC. We study the mechanism of action of viral RNAi suppressors and their role in virus evolution, host specificity and vector competence.

Schematic representation of the antiviral RNAi pathway. Figure adapted from Bronkhorst et al Fly 2013.
- piRNA pathway.
The PIWI-interacting pathway is a small RNA-mediated pathway, orginally described to be important for control of transposable elements in the germline. Interestingly, the piRNA pathway of Aedes mosquitoes has greated complexity compared to the pathway of genetic model organisms such as the fruit fly Drosophila melanogester. For example, Aedes aegypti encodes seven PIWI genes, which are expressed in both germline and somatic tissues, raising questions about their functions and mechanisms.
We found in collaboration with the Saleh lab (Pasteur Institute) that the piRNA pathway is involved in antiviral defense in mosquitoes. This pathway was previously known to be involved in control of transposable elements in germline tissues, but now seems to also process viral RNA substrates in Aedes mosquitoes. We study the non-canonical functions and mechanisms of the piRNA pathway in mosquitoes, such as antiviral defense and gene regulation.
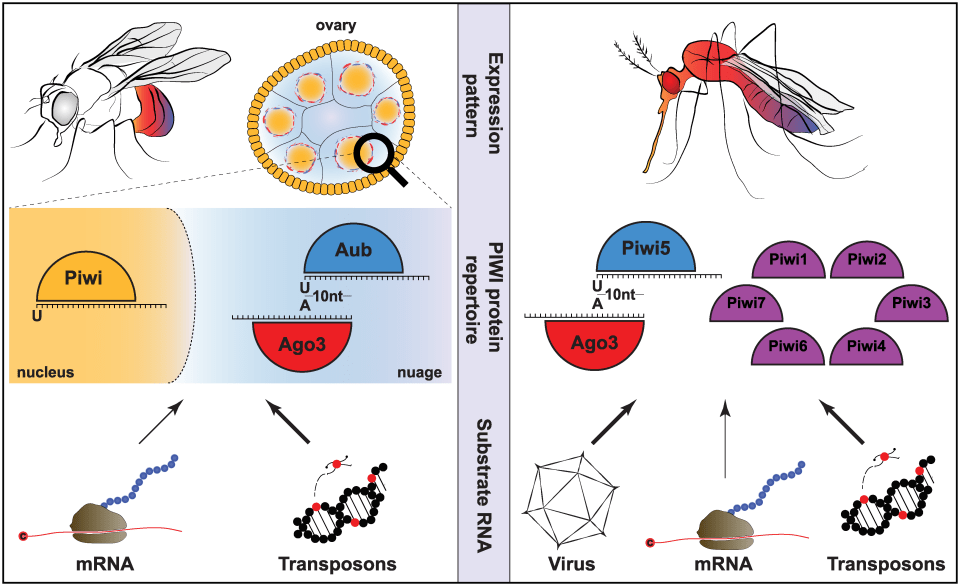
Key features of the piRNA pathway of Drosophila and Aedes mosquitoes. Figure from Miesen, Joosten & Van Rij, PLOS Pathogens 2016. https://doi.org/10.1371/journal.ppat.1006017.
- Innate antiviral immunity.
Vertebrates depend on a sophisticated network of interferon-based innate responses and adaptive immune responses for antiviral defense. Insects and other non-vertebrates, however, lack these responses, but – like all organisms – frequently encounter viral pathogens. Virus-infected mosquitoes often carry high viral loads, without overt pathology, suggesting that they have efficient mechanisms to tolerate or control infection. Understanding insect immunity is important from a fundamental perspective, but also central to our understanding of virus transmission by vector mosquitoes.
In addition to RNAi, antiviral immunity in insects relies on other, poorly characterized mechanisms. For example, the Jak-Stat pathway and the NF-kB pathways have been implicated in antiviral defense in some infection models, but how viral infection is sensed and the downstream antiviral effector genes remain undefined. Using a combination of transcriptomics and functional genomics in the fly, we wish to identify novel mechanism for antiviral defense and analyze epigenetic regulation of innate immunity.

Drosophila immune pathways. Figure adapted from Merkling et al J Insect Phys 2013
- Dengue virus-host interactions and antiviral compounds.
Despite the enormous global disease burden and health care costs associated with dengue and other arbovirus, no licensed drugs are available. Viruses rely on host factors for every step of their replication cycle and these host factors may be promising targets for therapeutic intervention. We recently screened a library of drug-like small molecules and identified a number of novel inhibitors of arbovirus replication. We study their mechanism of action and hope to identify the cellular target of the identified compounds. This work may yield leads for antiviral drug development, but will also provide insight into the viral replication cycle.
